Organisation
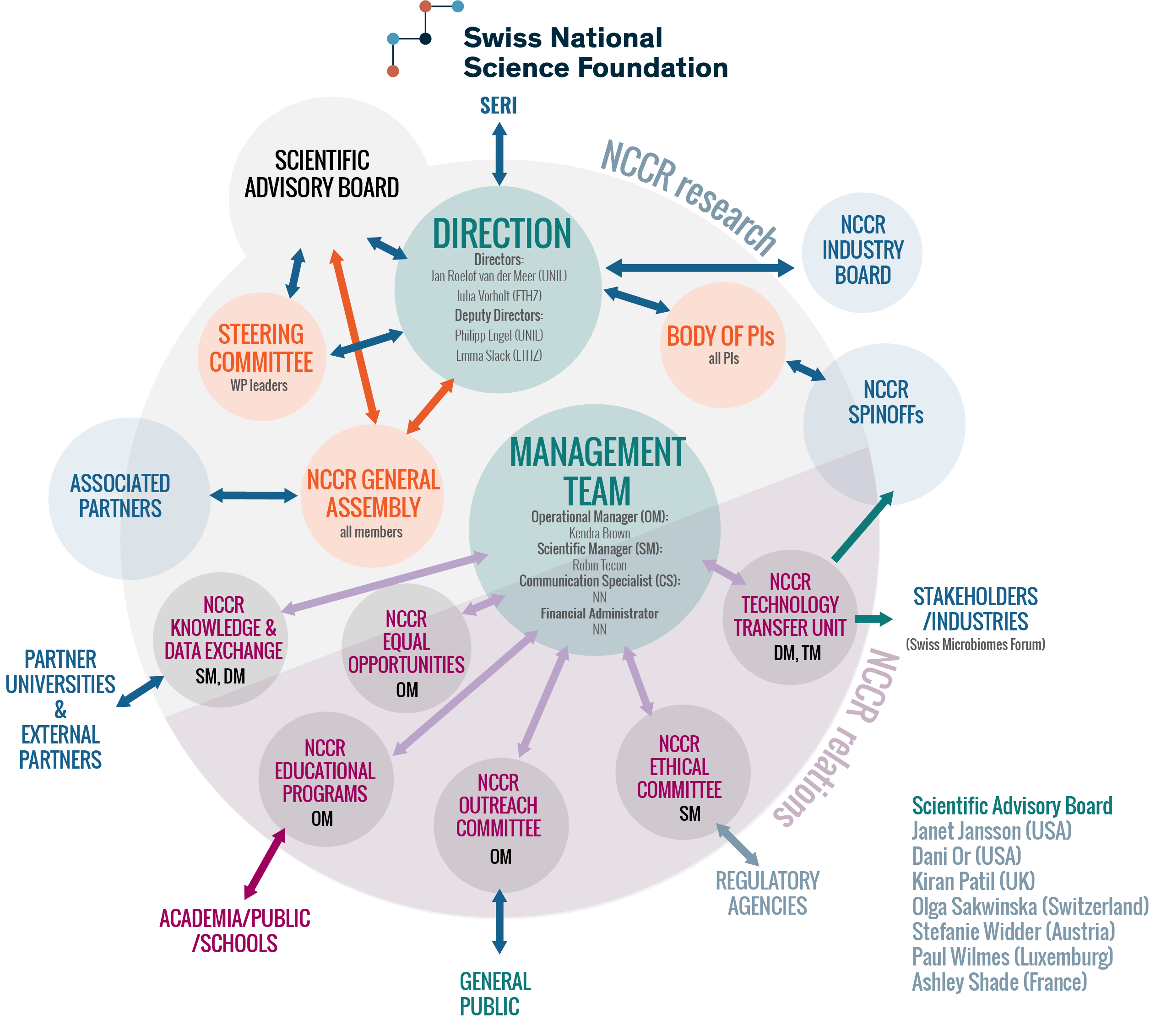
Management organigramme of the NCCR Microbiomes
Direction




Management team




Principal Investigators

























The NCCR Consortium
The NCCR Microbiomes is a consortium integrating 27 research groups from six institutions across Switzerland. The University of Lausanne (UNIL) acts as leading house, and ETH Zurich (ETHZ) as co-leading house. The NCCR is directed by Prof. Jan Roelof van der Meer (director, UNIL) and Prof. Julia Vorholt (co-director, ETHZ). Prof. Philipp Engel (UNIL) and Prof. Emma Slack (ETHZ) act as deputy director and co-director, respectively.
Five interlinked work packages (WP) tackle the research questions of the NCCR. Each WP gathers several research groups, with all groups being active across multiple WPs. One to two NCCR principal investigators (PIs) lead each WP and coordinate their efforts across the whole network.
The NCCR Microbiomes is managed by Dr. Robin Tecon (Programme Manager), Dr. Kendra Brown (Operational Manager), and Dr. Evelyne Schmid Osborne (Communication Specialist), who are in charge of the daily operations of the consortium, its scientific coordination, educational activities, knowledge and technology transfer, and equal opportunities.
The Steering Committee of the NCCR gathers the NCCR Direction, WP leaders and Operational and Programme managers. It coordinates research integration in close discussion with all Principal Investigators within the work packages, monitors research progress and difficulties, and advises on the use of shared or collective infrastructure.
A Scientific Advisory Board annually evaluates the NCCR, its research accomplishments, collaborations, knowledge and technology transfer, and education activities. The members of the Scientific Advisory Board are listed below.
All members of the NCCR Microbiomes constitute its General Assembly, which meets annually. The NCCR Ethical Committee is formed by PIs with translational research and clinical experience, and under the responsibility of the Programme Manager. It oversees the NCCR activities requiring coordination of ethical approval or more profound discussion of data and results dissemination or of societal implications of microbiome management.
Members of the NCCR Microbiomes Scientific Advisory Board
Associated Partners






NCCR Coworkers


















































































NCCR Alumni
























































